Research
African Integrative Evolutionary Genomics
We combine field work, laboratory work, and computational approaches to address fundamental questions about modern human evolutionary history and the genetic architecture of traits related to adaptation and disease risk in Africa. We are using an integrative genomics approach, incorporating genomic, proteomic, epigenetic, transcriptomic, metabolomic, and microbiome data obtained from ethnically diverse Africans living in distinct environments to identify genetic and environmental factors that play a role in a number of anthropometric, metabolomic, cardiovascular, and immune related traits (Figure 1).
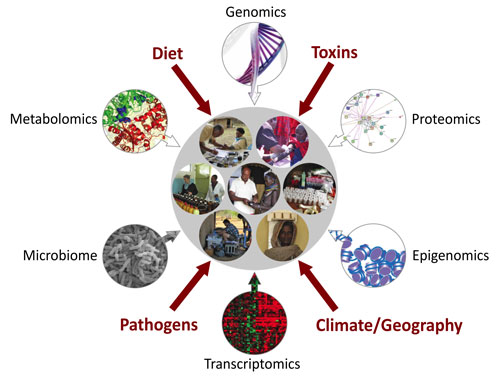
Figure 1. African Integrative Genomics approach to elucidate the genetic architecture of complex variable traits.
We incorporate genomic, proteomic, epigenetic, transcriptomic, metabolomic, and microbiome data obtained from ethnically diverse Africans living in distinct environments to identify genetic and environmental factors that play a role in a number of anthropometric, metabolic, cardiovascular, and immune related traits.
Many of these traits are likely to play a role in adaptation and some may play a role in disease susceptibility. We are characterizing gene and transcription networks to study how naturally occurring genetic and structural variation perturbs them. We are also studying how these networks are impacted by variable environmental factors such as lifestyle, diet, and infectious disease exposure. Additionally, we are examining patterns of genetic variation at the genome level among modern humans and non-human primates in order to elucidate the evolutionary forces (mutation, gene conversion/recombination, migration, drift, selection) that shape and maintain genetic variation in contemporary populations. These data are being used to reconstruct historical demographic and population differentiation events (including population expansion and contraction, subdivision, and migration) and to test hypotheses of modern human origins, including the possibility of introgression of archaic and modern human genomes.
African Genomic and Phenotypic Diversity Project
2191Most studies of human genomic variation and the genetic architecture of complex traits have focused on non-African populations. However, Africa is a critical region to study since it is the site of modern human origins, contains the greatest levels of human genetic variation (Figure 2), and is the source of the worldwide range expansion of modern humans in the past 100,000 years. Africa also has a high prevalence of several infectious diseases including HIV, malaria, and TB, resulting in millions of deaths per year. Additionally, several common complex diseases occur at higher frequency in African Americans, and are rapidly on the rise in urban regions of Africa, including hypertension, obesity, and type II diabetes. Differences in diet, climate, and exposure to pathogens among ethnically and geographically diverse African populations are likely to have produced distinct selection pressures, resulting in local genetic adaptations, some of which may play a role in disease susceptibility.
Because of differences in their demographic history and local adaptation, estimates of polygenetic risk scores fromEuropean-based studies may not be accurate in individuals of Africa ancestry, potentially resulting in inaccurate assessment of risk and lack of appropriate interventions. This European bias also impedes are ability to fully understand the genetic and environmental factors influencing risk of human disease and may exacerbate health disparities. Additionally, characterization of African genomic diversity is important for distinguishing pathogenic variants in clinical genomic studies since frequency of variants may differ among populations.
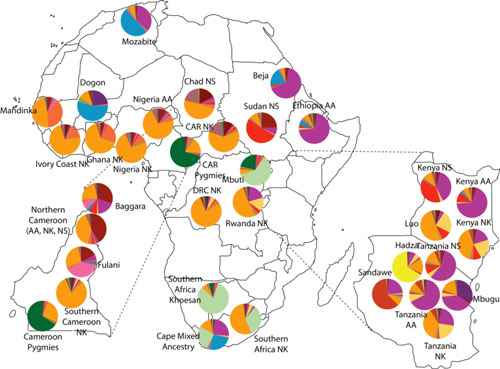
Figure 2: Patterns of genetic structure across African populations (from Tishkoff et al. Science 2009).
We inferred 14 genetically differentiated ancestral populations in Africa (indicated by colors)
The circles are colored according to the proportion of ancestry. Populations have been pooled by geographic regions and/or language classification. Our study of genome-wide variation in over 3200 Africans, the largest study of nuclear variation to date, demonstrates the extraordinarily high levels of genetic variation within and between populations in Africa, likely due to their unique demographic history as well as local adaptation to distinct environments.
Given the high levels of genetic diversity observed, even in small geographic regions, this study demonstrates the necessity to include a diverse set of Africans in human integrative genomic studies.
To address the disparity of human genomic studies in Africa, we and African collaborators have assembled an extensive and unique collection of DNA samples from >10,000 geographically and ethnically diverse Africans with distinct diets (hunter-gatherers, pastoralists, agriculturalists) and living in diverse environments with differing pathogen exposure (Figures 3). Additionally, for these samples we have detailed ethnographic and nutritional information as well as detailed phenotype data for a wide range of anthropometric, cardiovascular, metabolic, and immune related traits. Great care is taken to conduct this research in an ethical manner. We have undergone extensive ethical review and approval at the institutional and national level in each African country, obtained group and individual consent, and returned results to participants whenever possible. An additional goal is to help train African scientists and students and to help build resources within Africa for doing human genomics research.

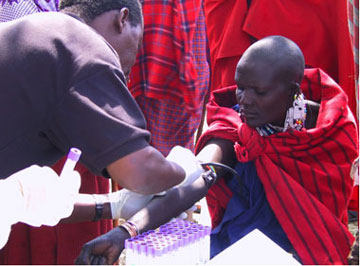




Figure 3. Field work in Africa.
In collaboration with our African colleagues, we are analyzing genomic data from this unique set of samples. From these studies, we will gain valuable knowledge of the genetic structure of African populations and the identification of markers that will be useful in gene mapping studies; we will learn about the correlation of environmental, cultural, linguistic, and genetic variation; we will be able to obtain estimates of demographic parameters and to test hypotheses of modern human origins and more recent population migration and differentiation events; we will identify functional regulatory elements that influence gene expression; we will identify genomic variants and pathways that play a role in human adaptation, phenotypic variation, and disease risk in African and African diaspora populations. Because non-African populations generally contain a subset of variation present in Africa, and because of low-levels of linkage disequilibrium in African populations, these studies will also be informative for fine-mapping of genetic variants associated with normal variable traits and disease risk in non-Africans.
KEY PUBLICATIONS
Gomez, F., J. Hirbo, and S.A. Tishkoff (2013) African population genetic history and evidence for local adaptation in African populations. Who are we? Human diversity and race from a contemporary genetics perspective. Chakravarti, A. (eds.) Cold Spring Harbor Press, In Press.
J. B. Hirbo, A. Ranciaro, and S. A. Tishkoff. (2013) Population structure and migration in Africa: correlations between archeological, linguistic, and genetic data. Causes and consequences of migration Benjamin Campbell and Michael H. Crawford (eds.). Cambridge University Press.
Lachance, J., B. Vernot, C.C. Elbers, B. Ferwerda, A. Froment, J.M. Bodo, G. Lema, W. Fu, T.B. Nyambo, T.R. Rebbeck, K. Zhang, J.M. Akey, S.A. Tishkoff (2012) Evolutionary history and adaptation from high-coverage whole-genome sequences of diverse African hunter-gatherers. Cell 150(3):457-469.
Williams, S.M and S.A. Tishkoff (2011) Exploring genomic studies in Africa. Genome Medicine 3(7):45.
Tishkoff, S. (2011) Interview with Sarah Tishkoff: perspectives for genetic research in African populations. Interview by Giovanni Destro-Bisol. Human Biology 83(5): 637-644.
Scheinfeldt, L. B., S. Soi, S.A. Tishkoff (2010) Colloquium paper: working toward a synthesis of archaeological, linguistic, and genetic data for inferring African population history. Proceedings of the National Academy of Sciences 107 (Suppl 2): 8931-8938.
Campbell, M.C., and S.A. Tishkoff (2010) The evolution of human genetic and phenotypic variation in Africa. Current Biology 20(4):R166-173.
Gonder, M.K., Locatelli, S., Ghobrial, L., Mitchell, M.W., Kujawski, J.T., Lankester, F.J., Stewart, C.-B., Tishkoff, S.A. (2011) Evidence from Cameroon reveals differences in the genetic structure and histories of chimpanzee populations. Proceedings of the National Academy of Sciences 108(12): 4766-4771.
Bryc, K., A. Auton, M.R. Nelson, J.R. Oksenberg, S.L., Hauser, S. Williams, A. Froment, J.M. Bodo, C. Wambebe, S.A. Tishkoff*, and C.D. Bustamante*(2010) Genome-wide patterns of population structure and admixture in West Africans and African Americans. Proceedings of the National Academy of Sciences 107(2):786-791. *Contributed equally as co-senior authors.
1bb7Tishkoff, S.A., F.A. Reed, F.R. Friedlaender, C. Ehret, A. Ranciaro, A. Froment, J.B. Hirbo, A.A. Awomoyi, J.M. Bodo, O. Doumbo, M. Ibrahim, A.T. Juma, M.J. Kotze, G. Lema, J.H. Moore, H. Mortensen, T.B. Nyambo, S.A. Omar, K. Powell, G.S. Pretorius, M.W. Smith, M.A. Thera, C. Wambebe, J.L. Weber, S.M. Williams (2009) The genetic structure and history of Africans and African Americans. Science 324(5930): 1035-1044.
Lambert, C.A., and S.A. Tishkoff (2009) Genetic structure in African populations: implications for human demographic history. Cold Spring Harbor symposia on quantitative biology 74: 395-402.
Campbell, M.C., Tishkoff, S.A. (2008) African genetic diversity: implications for human demographic history, modern human origins, and complex disease mapping. Annual Review Genomics and Human Genetics 9:403-433.
Sirugo, G., B.J. Hennig, A.A. Adeyemo, A. Matimba, M.J. Newport, M.E. Ibrahim, K.K. Ryckman, A. Tacconelli, R. Mariani-Costantini, G. Novelli, H. Soodyall, C.N. Rotimi, R.S. Ramesar, S.A. Tishkoff, S.M. Williams (2008) Genetic studies of African populations: an overview on disease susceptibility and response to vaccines and therapeutics. Human Genetics 123(6): 557-598.
Gonder, M.K., H.M. Mortensen, F.A. Reed, A. de Sousa, S.A. Tishkoff (2007) Whole-mtDNA genome sequence analysis of ancient African lineages. Molecular Biology and Evolution 24(3): 757-768.
Tishkoff, S.A, M.K. Gonder, B.M. Henn, H. Mortensen, N. Fernandopulle, C. Gignoux., A. Knight, P.A. Underhill, U. Ramakrishnan, F.A. Reed, J.L. Mountain (2007) History of click-speaking populations of Africa inferred from mtDNA and Y chromosome genetic variation. Molecular Biology and Evolution 24(10): 2180-2195.
Tishkoff, S.A. and M.K. Gonder. (2006) Human origins within and out of Africa. Anthropological Genetics M. Crawford (eds.). Cambridge University Press.
Reed, F.A. and S.A. Tishkoff (2006) African human diversity, origins, and migrations. Current Opinion in Genetics and Development 16(6):597-605.
Tishkoff, S A. and K.K. Kidd (2004) Implications of biogeography of human populations for ''race'' and medicine. Nature Genetics 36(11 Suppl): S21-27.
Tishkoff, S.A. and B.J. Verrelli (2003) Patterns of human genetic diversity: Implications for human evolutionary history and human disease. Annual Review Genomics and Human Genetics 4:293-340.
Tishkoff, S.A. and S.M. Williams. (2002) Genetic analysis of African populations: human evolution and complex disease. Nature Reviews Genetics 3(8):611-621.
Tishkoff, S.A., A. Goldman, F. Calafell, W.C. Speed, A.S. Deinard, B. Bonne-Tamir, J.R. Kidd, A.J. Pakstis,T. Jenkins, K.K. Kidd (1998) A global haplotype analysis of the myotonic dystrophy locus: implications for the evolution of modern humans and for the origin of myotonic dystrophy mutations. American Journal of Human Genetics 62(6):1389-1402.
Tishkoff, S.A., E. Dietzsch, W. Speed, A.J. Pakstis, K. Cheung, J.R. Kidd, B. Bonné-Tamir, A.S. Santachiara-Benerecetti, P. Moral, E. Watson, M. Krings, S. Pääbo, N. Risch, T. Jenkins, and K.K. Kidd (1996) Global patterns of linkage disequilibrium at the CD4 locus and modern human origins. Science 271(5254): 1380-1387.
Rawlings-Goss, Renata A., Michael C Campbell and Sarah A Tishkoff. Global Population-specific Variation in miRNA associated with Cancer Risk and Clinical Biomarkers. BMC Medical Genomics, Aug 28;7:53. doi: 10.1186/1755-8794-7-53. 2014
Campbell, M., Hirbo J., J. Townsend, S.A. Tishkoff. The Peopling of the African Continent and the Diaspora into the New World, Current Opinion in Genetics & Development, 15; 29C:120-132. 2014
Tishkoff, S.A. Strength in Small Numbers. Science, 349 (6254):1282-1283. 2015
Hansen ME, Hunt SC, Stone RC, Horvath K, Herbig U, Ranciaro A, Hirbo J, Beggs W, Reiner AP, Wilson JG, Kimura M, De Vivo I, Chen MM, Kark JD, Levy D, Nyambo T, Tishkoff SA,* Aviv A.*. Shorter telomere length in Europeans than in Africans due to polygenetic adaptation. Hum Mol Genet. Jun 1;25(11):2324-2330. 2016. *Co-Senior Author
Beltrame MH, Rubel MA, Tishkoff SA. Inferences of African evolutionary history from genomic data. Curr Opin Genet Dev. Dec;41:159-166. 2016
Yudell M, Roberts D, DeSalle R, Tishkoff S. Science and Society: Taking race out of human genetics, Science, Feb 5:351(6273):564-5. 2016
Kelly, D.E., Hansen, M.E.B, Tishkoff, S.A. Global variation in gene expression and the value of diverse sampling. Curr Opin Syst Biol. Feb;1:102-108. 2017
Matthew EB Hansen, Meagan A. Rubel, Aubrey G. Bailey, Alessia Ranciaro, Simon R. Thompson, Michael C. Campbell, William Beggs, Jaanki R. Dave, Gaonyadiwe G. Mokone, Sununguko Wata Mpoloka, Thomas Nyambo, Christian Abnet, Stephen J. Chanock, Frederic D. Bushman & Sarah A. Tishkoff. Population structure of human gut bacteria in a diverse cohort from rural Tanzania and Botswana. Genome Biology, Jan 22;20(1):16. doi: 10.1186. 2019
Sheinfeldt L , Sameer Soi, Charla Lambert, Wen-Ya Ko, Aoua Coulibaly, Alessia Ranciaro, Simon Thompson, Jibril Hirbo, William Beggs, Muntaser Ibrahim, Thomas Nyambo, Sabah Omar, Dawit Woldemeskel, Gurja Belay, Alain Froment, Junhyong Kim, Sarah Tishkoff. Genomic evidence for shared common ancestry of East African hunting-gathering populations and insights into local adaptation, Proc Natl Acad Sci U S A, Feb 19. pii: 201817678. doi: 10.1073/pnas.1817678116. [Epub ahead of print] PMID:30782801, 2019
Shaohua Fan, Derek E. Kelly, Marcia H. Beltrame, Matt EB. Hansen, Swapan Mallick, Thomas Nyambo, Sabah Omar, Dawit Wolde Meskel, Gurja Belay, Alain Froment, Nick Patterson, David Reich, Sarah A. Tishkoff. African evolutionary history inferred from whole genome sequence data, Genome Biology, 2Apr 26;20(1):82. 2019
Giorgio Sirugo, Scott M. Williams and Sarah A. Tishkoff, The missing diversity in human genetic studies, Cell, Mar 21;177(1):26-31. doi: 10.1016. 2019
The Genetic Basis of Resistance to Infectious Disease
244bIt is likely that infectious disease has played a major role in human evolution and in shaping genetic variation in the human genome. Thus, one focus of my laboratory is the study of human genetic variation and the evolutionary history of genes involved in resistance against infectious disease. We are characterizing patterns of genetic variation in candidate genes for resistance/susceptibility to malaria, TB, HIV, and trypanosomes in a set of globally diverse populations, but with an emphasis on African populations. Our goal is to identify functionally significant genetic variation that plays a role in susceptibility to infection. In addition, we are collaborating with laboratories that are studying genetic variation in the pathogens causing these disorders in order to examine co-evolution of infectious agents and their human hosts. Examples of this work include studies of G6PD, ICAM1, and glycophorin genes that play a role in malaria resistance and the APOL1 gene that plays a role in resistance to trypanosome infection, but is also associated with risk for kidney disease in African populations. We are also characterizing variation at the HLA and KIR loci in order to expand our knowledge of variation at these loci that play an important role in risk for disease and rejection of tissue and organ transplantation.
As part of the Chan Zuckerberg Institute funded Human Cell Atlas Consortium, we are generating an African Immune Cell Atlas. We are assessing normal transcriptomic and epigenetic variation of immune cells from ethnically diverse Africans using single cell RNA and ATAC sequencing. We are measuring individual and population-level variation in the cellular response to bacterial and viral immune stimulation. We are integrating that data with genomic sequencing data to identify genomic and epigenomic variation contributing to differences in immune response within and between populations, some of which may play a role in disease risk.
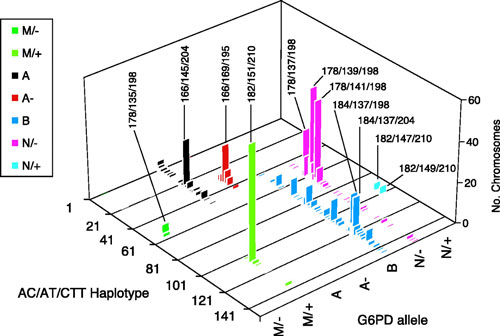
Figure 4.
(from Tishkoff, et al. Science, 2001)
Microsatellite haplotypes of chromosomes containing G6PD deficiency mutations (Mediterranean (M), A, and A-) as well as normal chromosomes (B and N). We see significantly higher linkage disequilibrium on the chromosomes containing G6PD deficiency mutations, consistent with a signature of natural selection, and we see that the African A and A- variants arose independently on a different chromosome background from the M variant, consistent with convergent evolution due to resistance from malaria infection.
KEY PUBLICATIONS
Ko, W.Y., P. Rajan, F. Gomez, L. Scheinfeldt, P. An, C.A. Winkler, A. Froment, T.B. Nyambo, S.A. Omar, C. Wambebe, A. Ranciaro, J.B. Hirbo, S.A. Tishkoff (2013) Identifying Darwinian selection acting on different human APOL1 variants among diverse African populations. American Journal of Human Genetics 93(1): 54-66.
Gomez, F., G. Tomas, W.Y. Ko, A. Ranciaro, A. Froment, M. Ibrahim, G. Lema, T.B. Nyambo, S.A. Omar, C. Wambebe, J.B. Hirbo, J. Rocha, S.A. Tishkoff (2013) Patterns of nucleotide and haplotype diversity at ICAM-1 across global human populations with varying levels of malaria exposure. Human Genetics 132(9):987-999.
Gomez, F., W.-Y. Ko, A. Davis, and S.A. Tishkoff. (2013) Impact of natural selection due to malarial disease on human genetic variation. Primates, pathogens and evolution. Developments in primatology: progress and prospects J.F. Brinkworth and K. Pechenkina (eds.) Springer publishing, New York. 38:117-160.
Ko, W.-Y., F. Gomez, and S.A. Tishkoff (2012) Rapid evolution of human erythrocyte-specific genes involved in malaria susceptibility. Evolution in the fast lane: Rapidly evolving genes and genetic systems. Rama S. Singh, Jianping Xu, and Rob J. Kulathinal (eds.). Oxford University Press.
Ko, W.Y., K.A. Kaercher, E. Giombini, P. Marcatili, A. Froment, M. Ibrahim, G. Lema, T.B. Nyambo, S.A. Omar, C. Wambebe, A. Ranciaro, J.B. Hirbo, S.A. Tishkoff (2011) Effects of natural selection and gene conversion on the evolution of human glycophorins coding for MNS blood polymorphisms in malaria-endemic African populations. American Journal of Human Genetics 88(6): 741-754.
Awomoyi A, G. Sirugo, M.J. Newport, and S.A. Tishkoff (2006) Global distribution of a novel trinucleotide microsatellite polymorphism (ATA)n in intron 8 of the SLC11A1 gene and susceptibility to pulmonary Tuberculosis. European Journal of Immunogenetics 33(1):11-15.
Tarazona-Santos, E., S.A. Tishkoff (2005) Divergent patterns of linkage disequilibrium and haplotype structure across global populations at the interleukin-13 (IL13) locus. Genes and Immunity 6(1):53-65.
Tishkoff, S.A. and B.J. Verrelli (2004) G6PD deficiency and malarial resistance in humans: insights from evolutionary genetic analyses Evolutionary Aspects of Infectious Disease. K. Dronamraju (eds.) Cambridge University Press, 2004.
Verrelli, B.C., J.H. McDonald, G. Argyropoulos, G. Destro-Bisol, A. Froment, A. Drousiotou, G. Lefranc, A.N. Helal, J. Loiselet, S.A. Tishkoff (2002) Evidence for balancing selection from nucleotide sequence analyses of human G6PD. American Journal of Human Genetics 71(5):1112-1128.
Tishkoff, S.A., R. Varkonyi, N. Cahinhinan, S. Abbes, G. Argyropoulos, G. Destro-Bisol, A. Drousiotou, B. Dangerfield, G. Lefranc, J. Loiselet, A. Piro, M. Stoneking, A. Tagarelli, G. Tagarelli, E.H. Touma, S.M. Williams, A.G. Clark (2001) Haplotype diversity and linkage disequilibrium at human G6PD: recent origin of alleles that confer malarial resistance. Science 293(5529):455-462.
The Genetic Basis of Adaptation in Humans
We currently know little about how changes at the genetic level correlate with phenotypic changes and adaptation to novel environments during recent human evolutionary history. Additionally, it has been hypothesized that genetic mutations associated with common complex diseases (e.g. hypertension, diabetes, obesity, asthma, arthritis, allergies, etc.) may be at high frequency in modern populations because they were adaptive in ancient environments. Thus, characterization of signatures of natural selection in genes that are of adaptive significance may be of use for identifying functionally significant variants, some of which may play a role in human disease. We are particularly interested in identifying local adaptation in culturally and geographically diverse Africans, because of the possibility that selective forces, and genetic variants, may be geographically restricted. We are also interested in developing and applying methods for distinguishing balancing selection, soft selective sweeps (i.e. selection from standing variation), and selection of loci involved in complex traits. Recent examples of our studies of local adaptation in Africa includes the identification and characterization of natural selection at loci that play a role in lactose tolerance in African pastoralists (Figure 5), the genetic basis of adaptation to high altitude in Ethiopia, the genetic basis of the short stature trait in Western African Pygmies, and the genetic basis of skin pigmentation in Africa (in addition to the studies of adaptation at genes involved in infectious disease resistance described above).
We are particularly interested in understanding how natural selection has shaped regulation of gene expression. The majority of candidate loci from GWAS and selection scans are in non-coding regions of the genome. We are using massively parallel reporter assays to identify enhancer activity of thousands of cis-regulatory elements identified in GWAS and selection screens. Candidate genes regulated by these enhancers are identified using chromatin capture-based techniques and the impact of candidate regulatory variants on gene expression and phenotypes of interest are validated using CRISPR/Cas9 editing in vitro and/or in vivo
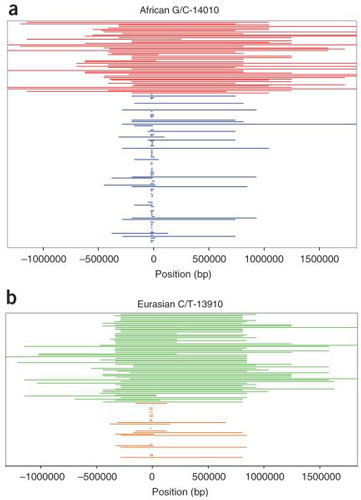
Figure 5. Comparison of tracts of homozygous genotypes flanking the lactase persistence–associated SNPs.
(from Tishkoff et al., Nature Genetics, 2007)
(a) Kenyan and Tanzanian C-14010 lactase-persistent (red) and non-persistent G-14010 (blue) homozygosity tracts. (b) European and Asian T-13910 lactase-persistent (green) and C-13910 non-persistent (orange) homozygosity tracts. The extended haplotype homozygosity on chromosomes with the lactase persistence associated mutations indicates very recent and strong positive selection of these mutations in populations that practice cattle domestication and dairying.
KEY PUBLICATIONS:
Campbell, M.C., A. Ranciaro, D. Zinshteyn, R. Rawlings-Goss, J.B. Hirbo, S.I. Thompson, D. Woldemeskel, A. Froment, J.B. Rucker, S.A. Omar, J.-M. Bodo, T. Nyambo, G. Belay, D. Drayna, P.A.S. Breslin, S.A. Tishkoff (2013) Origin and differential selection of allelic variation at TAS2R16 associated with salicin bitter taste sensitivity in Africa. Molecular Biology and Evolution, Oct 30. [Epub ahead of print].
Jarvis, J.P., L.B. Scheinfeldt, S. Soi, C. Lambert, L. Omberg, B. Ferwerda, A. Froment, J.M. Bodo, W. Beggs, G. Hoffman, J. Mezey, S.A. Tishkoff (2012) Patterns of ancestry, signatures of natural selection, and genetic association with stature in Western African pygmies. PLoS Genetics 8(4): e1002641.
Scheinfeldt, L.B., S. Soi, S. Thompson, A. Ranciaro, D. Woldemeskel, W. Beggs, C. Lambert, J.P. Jarvis, D. Abate, G. Belay, S.A. Tishkoff (2012) Genetic adaptation to high altitude in the Ethiopian highlands. Genome Biology 13(1):R1.
Mortensen, H., A. Froment, G. Lema, J.M. Bodo, M. Ibrahim, T.B. Nyambo, S.A. Omar, S.A. Tishkoff (2011) Characterization of genetic variation and natural selection at the arylamine N-Acetyltransferase (NAT) genes in global human populations. Pharmacogenomics 12(11):1545-1558.
Scheinfeldt, L.B. and S.A. Tishkoff (2010) Living the high life: high-altitude adaptation. Genome Biology 11(9):133.
Tishkoff, S. A., F. A. Reed, A. Ranciaro, B.F. Voight, C.C. Babbitt, J.S. Silverman, K. Powell, H.M. Mortensen, J.B. Hirbo, M. Osman, M. Ibrahim, S.A. Omar, G. Lema, T.B. Nyambo, J. Ghori, S. Bumpstead, J.K. Pritchard, G.A. Wray, P. Deloukas (2007) Convergent adaptation of human lactase persistence in Africa and Europe. Nature Genetics 39(1):31-40.
Verrelli, B.J. and S.A. Tishkoff (2004) Signatures of selection and gene conversion associated with human color vision variation. American Journal of Human Genetics 75(3):363-375.
Joseph Lachance and Sarah A. Tishkoff. Population Genomics of Human Adaptation. Annual Review of Ecology, Evolution, and Systematics, 44:123-143, 2013
Scheinfeldt, L. B., Tishkoff, S. A. Recent human adaptation: genomic approaches, interpretation and insights. Nature reviews. Genetics 14(10): 692-702, 2013
Gomez, F., J. Hirbo, and S.A. Tishkoff. African Population Genetic History and Evidence for Local Adaptation in African Populations. Who Are We? Human Diversity and Race from a Contemporary Genetics Perspective. Chakravarti, A. (eds.). Cold Spring Harbor Press, 2014
Ranciaro, A., Campbell, M.C., Hirbo, J.B., Ko, W.-Y., Froment, A., Destro-Bisol, G., Kotze, M.J., Ibrahim, M., Nyambo, T., Omar, S.A., Tishkoff, S.A. Genetic origins of lactase persistence and the spread of pastoralism in Africa, American Journal of Human Genetics, 3;94(4):496-510, 2014
Campbell MC, Ranciaro A, Zinshteyn D, Rawlings-Goss R, Hirbo J, Thompson S, Woldemeskel D, Froment A, Omar SA, Bodo JM, Nyambo T, Belay G, Drayna D, Breslin PA, Tishkoff SA. Limited evidence for adaptive evolution and functional effect of allelic variation at rs702424 in the promoter of the TAS2R16 bitter taste receptor gene in Africa. J Hum Genet. Jun;59(6):349-52. 2014
Fan S, Hansen ME, Lo Y, Tishkoff SA. Going global by adapting local: A review of recent human adaptation. Science. Oct 7;354(6308):54-59. 2016


