Project 4: Animal Models of Mesothelioma
Significance
Malignant mesothelioma (MM) is a highly aggressive, treatment-unresponsive cancer usually caused by exposure to asbestos such as that found in Ambler, Pennsylvania. With estimates of >20 million individuals at risk worldwide, new approaches in disease management and prevention are badly needed. The genetic basis for MM has historically focused on somatic mutations of the tumor suppressor genes CDKN2A and NF2 as key alterations influencing initiation and progression.
Recently, the BAP1 ubiquitin carboxy-terminal hydrolase, has been strongly implicated as a major player in MM based on genetic analyses. Germline BAP1 mutations were found in families with a high incidence of MM and other cancers, and somatic BAP1 alterations occurred in MMs, consistent with biallelic inactivation of a tumor suppressor. Moreover, somatic BAP1 mutations are common in sporadic MMs, often in combination with alterations of NF2 and CDKN2A. The genetic and biochemical mechanisms by which BAP1 mutations predispose to MM and how BAP1 interacts genetically with CDKN2A and NF2 to influence MM pathology and therapeutic response are largely unknown. Genetically engineered mouse models can be extremely useful in advancing the understanding tumor development.
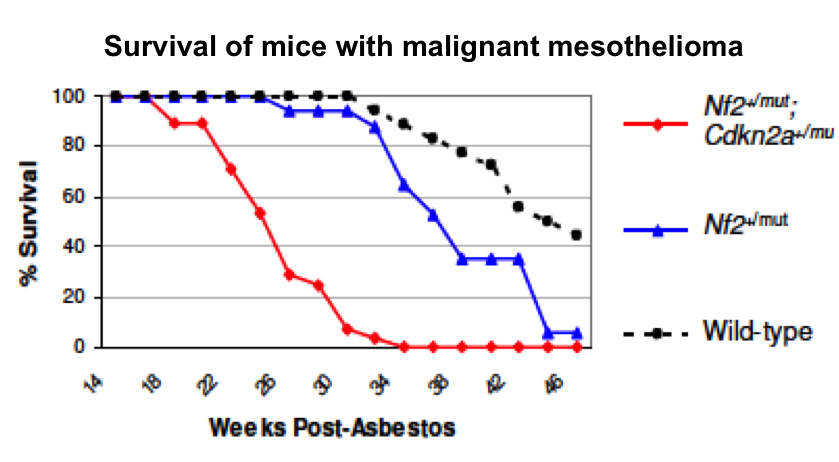
Previous in vivo carcinogenicity studies of crocidolite, asbestos of the amphibole type, have revealed that mouse models with heterozygous mutations (+/mut) of either Nf2 or Cdkn2a exhibit accelerated induction of MM compared to wild-type (+/+) mice. Moreover, crocidolite-exposed mice with mutations of both Nf2 and Cdkn2a (Nf2+/mut;Cdkn2a+/mut) showed further acceleration of MM induction and a more aggressive tumor phenotype, strongly supporting the notion that inactivation of multiple tumor suppressor genes can cooperate to drive MM pathogenesis. However, whether specific epigenetic alterations are also required for MM development is ill-defined. Moreover, whether Nf2+/mut;Cdkn2a+/mut mice are similarly vulnerable to other forms of asbestos, such as tremolite or chrysotile, and whether Bap1+/mut mice are predisposed to the effects of asbestos, are currently unknown. Additionally, whether biological remediation of asbestos abolishes its carcinogenicity in vivo has not been formally tested in such relevant mouse models of MM.
The Testa and Simmons Labs have joined forces to pursue the following Specific Aims:
- Use a direct in vivo genetic approach to determine if both Bap1+/mut mice and Nf2+/mut;Cdkn2a+/mut mice are predisposed to the induction of MM by both tremolite and chrysotile.
- Use Nf2+/mut;Cdkn2a+/mut mice to ascertain whether remediation of asbestos suppresses its carcinogenic potential.
- Identify genome-wide epigenetic modifications resulting in changes in expression, which in turn are associated with MM formation and progression.
The proposed studies by two Co-PIs with complementary expertise in genetics and epigenetics represent a comprehensive approach to yield novel basic insights into asbestos carcinogenicity and mechanisms that drive MM development and dissemination, with translational implications for understanding tumor susceptibility and prevention. Project 4 will utilize remediated asbestos from Project 1 to test directly, in genetic model systems, if biological remediation of asbestos abrogates its carcinogenic potential in vivo. It will also provide animal models and expertise for the chemoprevention studies proposed in Project 5 and take advantage of the biomarker work in Project 6 to confirm that remediated fibers are less toxic. Insights from this project are also expected to provide rationale for developing asbestos remediation and mitigation strategies that will improve risk management (Projects 1 and 2).
Project 4 will interact with the Administrative Core for overall direction and to facilitate progress. It will interact with the Community Engagement Core to describe the project in layperson terms to the Stakeholder Advisory Group in Ambler on an annual basis. It will inform the Research Translation Core about progress towards the identification of biomarkers of disease onset that could be validated in humans and oncogenic pathways for targeted therapies. The Biostatistics Support Research Core will be used for study design and analysis of data. Pre- and post-doctoral fellows will participate in the Interdisciplinary Research Training Core.
Project Leaders
Joseph R. Testa, PhD, Project Leader
Senior Member, Population Science Division, Fox Chase Cancer Center
Joseph.Testa@fccc.edu
Rebecca A. Simmons, MD, Co-Project Leader
Hallam Hurt Professor in Neonatology, Department of Pediatrics, Perelman School of Medicine, University of Pennsylvania
rsimmons@mail.med.upenn.edu
Publications
Click Here to view all publications