Research Interests
The Kaestner lab employs modern mouse genetic approaches, such as gene targeting, tissue-specific and inducible gene ablation, to understand the molecular mechanisms of organogenesis and physiology of the liver, pancreas and gastrointestinal tract. We also use next-generation sequencing to look at differences between the transcriptome and epigenome of normal vs diseased tissues.
Some of our current research includes:
The mammalian gut epithelium is a highly organized and dynamic system which requires continuous controlled proliferation and differentiation throughout life. Proliferation, cell migration and cell adhesion all must be tightly controlled in order to prevent either inflammatory diseases or epithelial cancers. As with many other vertebrate organs, the digestive tract develops from heterogeneous embryonic origins. While the musculature and the connective tissue are derived from lateral plate mesoderm, the epithelium is derived from the endoderm. We have identified a novel member of the winged helix gene family termed Foxl1 which is expressed in the gut mesoderm and have begun its functional analysis in vivo through targeted mutagenesis in mice. Null mutations in the mesodermal transcription factor Foxl1 result in dramatic alterations in endoderm development, including epithelial hyperproliferation. We have now identified APC/Min and GKLF as downstream targets of Foxl1 and have begun the analysis of these genes in gastrointestinal differentiation by tissue-specific gene ablation.
Recent studies including our own suggest a marked increase in β-cells expressing key components of cellular senescence in islets from Type 1 diabetes (T1D) and Type 2 diabetic (T2D) patients, implicating β-cell senescence as a critical contributor to islet dysfunction. Recently, it was reported that ablation of senescent cells by non-specific senolysis in mouse models of T1D and T2D improved disease outcome. However, these studies did not determine which senescent cell type was relevant to the beneficial effect, nor did they address to what extent senescence occurs in the human endocrine pancreas before and during the development of T1D and T2D. We are working to determine the prevalence, transcription signatures, and epigenomic landscapes of β-cell senescence in T1D, pre-T1D and T2D donors using immunostaining, imaging mass cytometry, single cell RNAseq, single cell ATACseq, DNA methylome determination and Cut-and-Tag analysis for key histone marks. In addition, we are testing the hypothesis that irreparable damage to telomeres drives senescence in β-cells, a possible scenario that could provide a mechanism for senescence to occur in islet cells of diabetic patients. We are further seeking to uncover whether metabolic and/or inflammatory stressors drive senescence in human β-cells and employing a novel transgenic mouse, the ‘SenKiller’ model, to enable cell-type specific and inducible ablation of senescent cells in any lineage including β-cells. Together, these studies will provide a definitive answer to the question if senescent β-cells are critical in the development of glucose intolerance in models of T2D and islet autoimmunity in models of T1D.
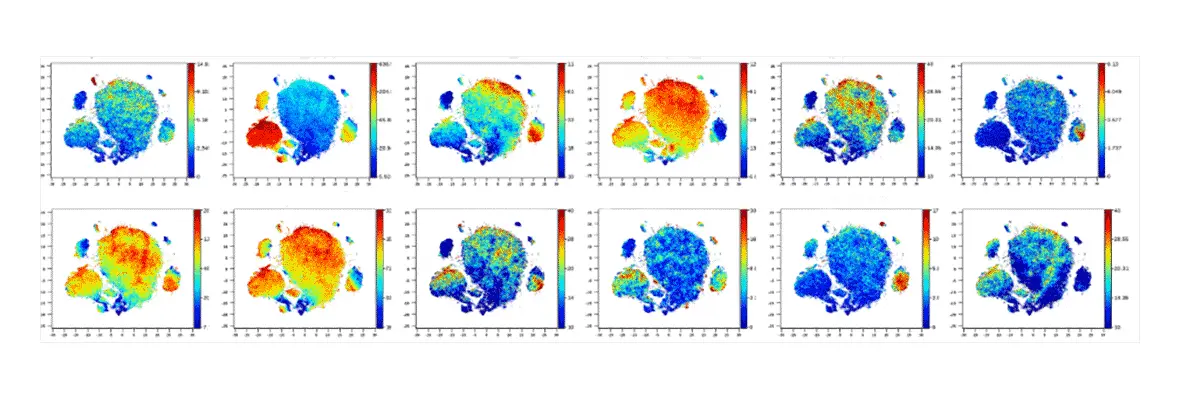
The Human Pancreas Analysis Program (HPAP) is a recently established National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) initiative that consists of two related programs studying both type 1 and type 2 diabetes, HPAP-T1D and HPAP-T2D. The programs are designed to define the molecular pathogenesis of islet dysfunction by studying human pancreatic tissue samples from organ donors with no history of diabetes, or at varying stages of T1D and T2D progression. HPAP generates detailed datasets of physiological, histological, transcriptomic, epigenomic, and genomic information using state of the art techniques. Importantly, all data collected, generated, and analyzed by HPAP-T2D are made immediately and freely available through a centralized database, PANC-DB, thus providing a comprehensive data resource for the diabetes research community.
More information can be found on PANC-DB.