The mission of the Molecular and Translational Immunotechnology Core is to further innovative, interdisciplinary and translational research that enhances our understanding of the pathogenesis and immunopathogenesis of HIV/AIDS; provides new approaches toward understanding cellular, humoral and innate responses to HIV; and develops novel HIV therapy and vaccine strategies. To achieve these goals, the Core provides state-of-the-art immunological services and reagents; specialized technology; leadership, expertise and advice; and collaborative support in the area of immunological research to the Penn CFAR community. The Core also works closely with the Perelman School of Medicine Human Immunology Core (HIC).

Available Services
The Molecular and Translational Immunotechnology Core provides a wide range of services and reagents to benefit the immunological research of the Penn CFAR Community. Services are provided by facilities in the University of Pennsylvania. Services include:
- Primary Cell Purification (Riley Lab, 8th Floor Smilow Center for Translational Research)
- Freshly isolated primary PBMCs, CD4, CD8, T cells, NK, monocytes and B cells
- CCR5 deficient T cells and monocytes from Δ32 donors
- Sample processing and immunology assays (HIC, 410, 411 and 412 Stellar Chance Labs)
- Specimen Processing, Banking, & Sample Shipment
- Luminex assays- multiplex bead arrays (FlexMAP3D)
- Flow Cytometry- standard and customized assays
- ELISPOT
- ELISA
- Digital ELISA (Simoa)- ultra-sensitive molecular detection
- Sequencing of Human & Murine IgH rearrangements
- Sequencing of Human TCR Vβ rearrangements
- Data analysis pipeline for repertoire studies
- Customized Data Analysis- in-house data QC, analysis and visualization methods for immune repertoire profiling
- Scientific Consultation and Assay Development
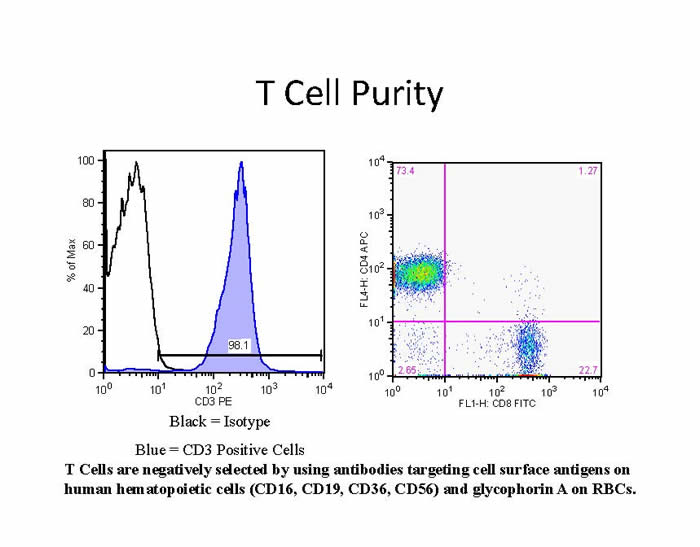
Within the last several years there has been a greatly increased demand for primary cells in both HIV and immunological research. There are multiple reasons for this. There has been a shift in the field from the use of cell lines to primary cells for research. For example, during the transition from primary cells to transformed cell lines, critical signal transduction pathways are altered, which complicates the extrapolation of data to primary cells. Moreover, certain cell types, such as dendritic cells (DC), do not have cell lines that faithfully model their activities, necessitating the use of primary cells. Lastly, certain experiments require cells with a particular genotype or HIV status. Transformed cell lines are popular because they are inexpensive, require little expertise to use, and are easy to propagate; the major goal of this Immunology Core service is to make primary cells as accessible and easy to use as transformed cells.
For the most part, investigators prefer cells that have been purified via a negative selection process, which leaves them as minimally manipulated as possible. Cells are obtained from healthy donors by magnetic. Bead-based selection developed by STEMCELL Technologies. Cells are generally provided fresh to investigators or, for those who require large batches of single-donor cells to be used at different time points, can be provided as frozen aliquots. The cell purity is always greater than 90% and is routinely greater than 95%. During the time the MTIC has served the Penn CFAR community, we have isolated cells from over 600 apheresis products; a sample summary for one such product is shown in Table 1. In addition, purified cells from HIV-1 infected donors are available upon request.
Table 1. Typical Cell Request
| Cell Type | # Requested | Purity |
|---|---|---|
| CD4 T cell | 350 million | 95% |
| CD8 T cell | 100 million | 97% |
| Monocytes | 350 million | 95% |
| PBMC | 200 million | N/A |
| CD8-depleted PMC | 100 million | 98% |
This is an ideal core function because the effort required to isolate 10 million cells is the same as the effort to isolate one billion cells. Rather than having all laboratories who wish to work with primary cells obtain their own IRB-approved protocol, recruit donors, and develop the expertise to purify cells, the MTIC performs all of these functions, and since the entire leukapheresis product is utilized, the price per cell is much less than if the researchers were to obtain their own cells. A major part of the appeal and popularity of this service is that it removes the regulatory burden required to work with primary cells: The MTIC maintains the IRB-approved protocol and the individual core users receipt of these cells is considered secondary use of de-identified human specimens which is not considered human use by both the NIH and our IRB.
Donors are not tested for any infectious agents. So as with all blood borne products, users are encouraged to follow universal precautions.
Ordering information:
Order cells at the Human Immunology Core
Primary Contact: James L. Riley, Ph.D. | rileyj@upenn.edu | 215-573-6792
- Specimen Processing, Banking, & Sample Shipment– The MTIC leverages the existing infrastructure and expertise of staff in the Human Immunology Core to perform sample processing, cryopreservation, banking and shipping services in support of early-phase clinical trials. Services include processing blood, biopsies and cadaver tissues as well as biological fluids (serum, plasma, CSF etc.). The core has dozens of standard operating procedures (SOPs) for sample processing, biobanking and shipping. Staff are internally and externally qualified to perform these procedures and customized workflows are available. At the beginning of a study, clients meet with core lab staff to customize sample processing, storage and biobanking workflows as well as metadata capture from samples. The Core has a laboratory information management system (LIMS, LabVantage) and can interface with other data entry/collection systems, as needed for a specific study. On-site storage is available on a limited basis. Core lab freezers are subject to regular monitoring including electronic monitoring.
- Luminex assays. The Core has two FlexMAP3D instruments for multiplex bead array analysis. Assays typically use commercially available kits, but homebrew assays are also possible. Typical analytes include cytokines, autoantibodies, anti-viral antibodies etc. The Core has excellent relationships with vendors and obtains discounts on assay kits that help reduce the cost to CFAR investigators.
- Flow Cytometry. Multicolor immunophenotyping assays are available. The Core has standard panels for the analysis of circulating immune cell subsets including B cells, plasma cells, T cells, NK cells and monocytes. Customized assays of antigen-enriched or other specific immune cells in blood, bone marrow, fluid or tissue samples are also available. The Core has also developed a RBC immunophenotyping panel for enumeration of fetal RBCs and evaluation of RBC maturation.
- ELISPOT. The Core has an ImmunoSpot reader and can perform and/or read ELISPOT assays on a fee-for-service basis. ELISPOT assays can be used to quantify the frequency and function of antigen-enriched T cells or B cells by measuring cytokine or antibody production by activated cells.
- ELISA. The Core performs enzyme linked immunosorbent assays to measure antibodies or antigens of interest, usually employing commercially available kits. The Core has a CLARIOstar multimodal microplate reader, enabling read-outs using a variety of formats, e.g., fluorescence, optical density etc.
- Digital ELISA. The Core has a Quanterix HD-X analyzer for ultra-sensitive detection of biomarkers, leveraging Simoa bead technology, which offers up to 1000-fold greater sensitivity than traditional immunoassays. Singleplex, multiplex and homebrew assays are possible on this platform.
- Immune repertoire profiling. Immune repertoires are analyzed by next-generation sequencing of antibody (a.k.a. BCR, Ig) and T cell receptor (TCR) gene rearrangements. Immune repertoires are helpful for understanding immune responses, tissue immunity, making disease group comparisons or performing longitudinal monitoring (e.g., minimal residual disease, therapeutic effects etc.) Human BCR/TCR and murine (C57Bl/6 and BALB/c) BCR sequencing are available. Bulk sequencing is available using DNA or RNA-based samples. Unique molecular identifiers are available for RNA-based sequencing projects that utilize high sequencing depth on moderate template numbers for the creation of high-fidelity consensus reads. For human BCR sequencing, we provide primers that can amplify full-length (leader region), near full-length (FR1) or short-read (FR3) are available. For RNA-based sequencing, antibody isotyping is available with different constant region primers. Single cell sequencing is available in conjunction with other Cores.
- Data analysis. The Core provides routine data QC and processing and have an in-house data analysis platform for immune repertoire profiling data (ImmuneDB), with publicly shared analysis methods and code (available on GitHub). The Core provides clients with data annotation and can assist with data sharing using the Adaptive Immune Receptor Repertoire (AIRR) Community miAIRR standard on a fee-for-service basis. All immune repertoire profiling data that are generated for core lab clients are available, including raw fastq files, if requested by the investigator. The Core also provides project-specific data analysis and visualizations that are fully customizable for clonal landscape, V gene usage, public clones, CDR3 edit distances, isotyping, clone tracking, diversity, somatic mutation, selection and clonal lineage analysis etc..
- Scientific consultation and assay development. Core leadership provides consultation on projects, protocols, reagents, assays and new technology. We provide letters of collaboration for grant proposals as well as detailed budgets for projects and proposal planning.
Ordering Information
Please visit the HIC website for assays, pricing and ordering information.
Contacts
Nina Luning Prak, MD, PhD | luning@pennmedicine.upenn.edu | 215-746-5768 for general inquiries, new technology, letters of collaboration, scientific consultation
Honghong Sun, PhD | hsun2@pennmedicine.upenn.edu | 215-573-5367 for sample processing, clinical trial workflows, shipping, inventory, cellular immunology assays (flow cytometry, Luminex, ELISA, digital ELISA, ELISPOT)
Wenzhao Meng, PhD | wmeng@pennmedicine.upenn.edu | 215-573-5367 for molecular immunology assays including antibody and T cell receptor gene rearrangement analysis

